Research
We are a genomics and bioinformatics research group at Zhejiang University Liangzhu Laboratory in Hangzhou, China. Our research combines 'wet-lab' experiments, computational meta-data analysis, and machine learning algorithms. Our research areas are:
1. Computational algorithm and bioinformatics tool development to guide experimental design or make better use of different types of omics data.
We are fascinated by the omics view of life. As sequencing technologies quickly advance, we are facing with the challenge of interpreting omics big data. One of the major efforts in the lab is to develop computational algorithms and bioinformatics tools customized for both the underlying biology and data type of interest. We have developed methods for high precision detection of somatic mutations directly from scRNA-seq data, and highly sensitive gene fusion detection algorithms that outperforms the targeted sequencing approach. We are amazed by how much room of improvement there is and how badly such methods and tools are needed.
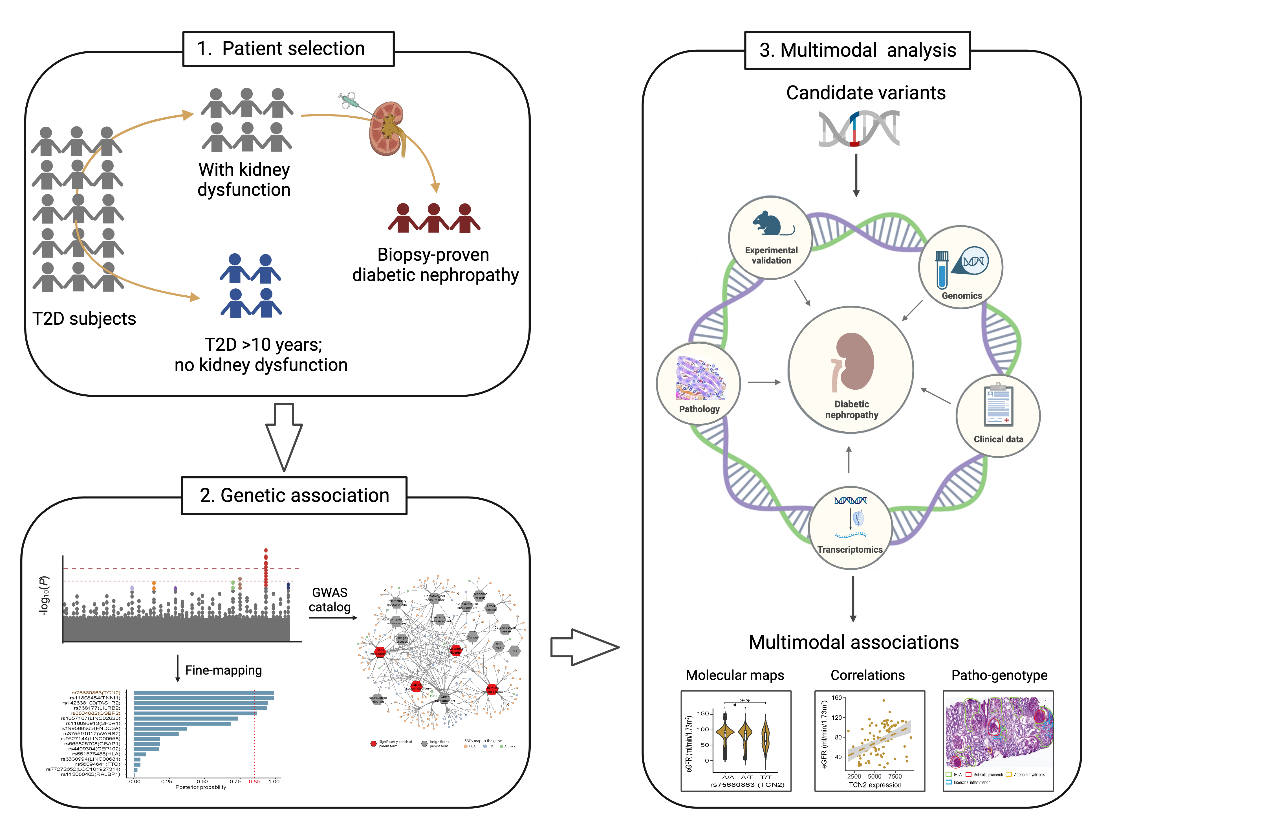
2. Multi-omic data analysis and studies of important diseases like diabetic nephropathy.
Working in an interdisciplinary field, we help solve essential biological and translational questions by collaborating closely with others to conduct multi-omics analysis and studies. Examples of diseases that we are working on include diabetic nephropathy and african swine flue. We are eager to learn from other experts as well as making our impact through such collaborative work.
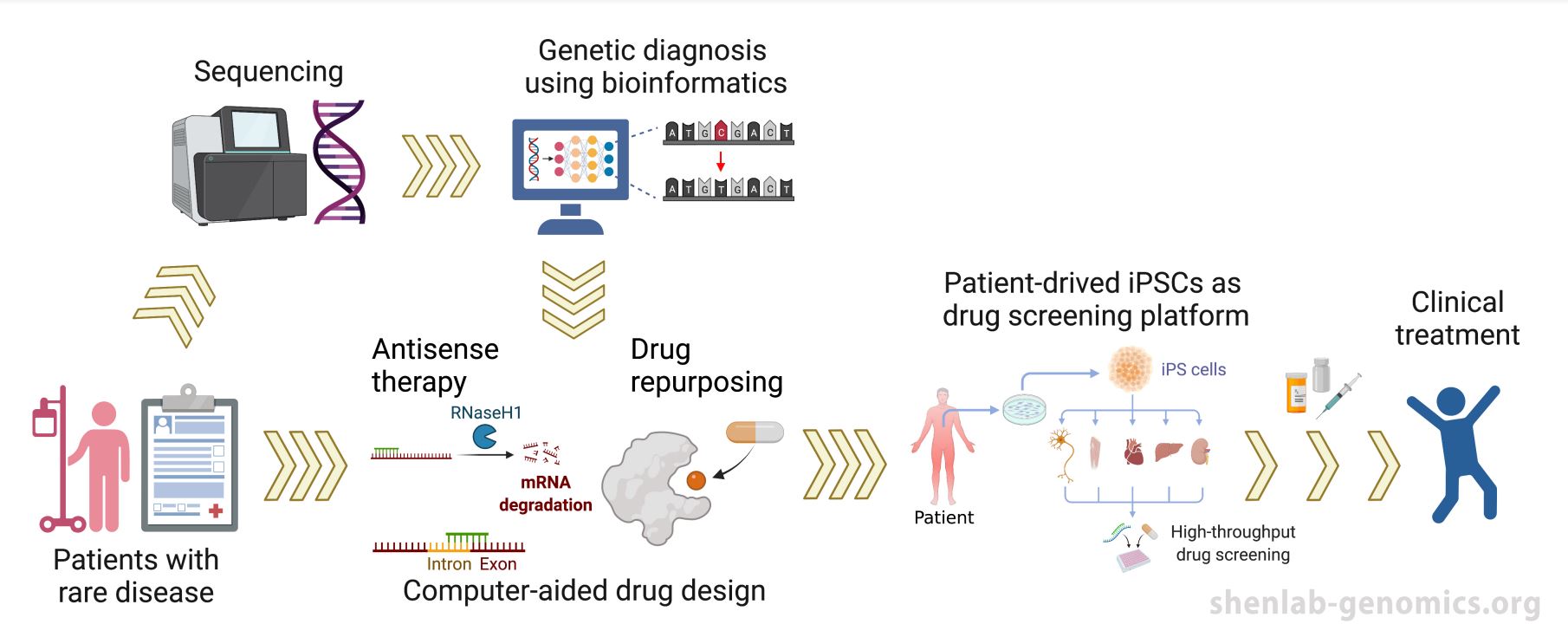
3. Computational guided diagnosis and drug development for Rare Genetic diseases.
We are committed to the precision medicine for rare genetic disease, both in terms of precise diagnosis and the development of therapeutic strategies. Built upon close collaboration with several hospitals, we are continuing to collect sequencing data of rare diseases patients, and to develop a series of algorithms to improve the genetic diagnosis. Meantime, we have built in-house iPSC differentiation protocols as our drug screening platform. Based on our computationally guided drug design, we firmly believe that fast and precise diagnosis and therapeutics that is customized for individual patients, will be achievable.